Technology peripherals
Technology peripherals
 AI
AI
 Efficiently and accurately predict DDI, the explanatory drug AI model of Fuzhou University and Yuanxing Intelligent Drug Team was published in Nature sub-journal
Efficiently and accurately predict DDI, the explanatory drug AI model of Fuzhou University and Yuanxing Intelligent Drug Team was published in Nature sub-journal
Efficiently and accurately predict DDI, the explanatory drug AI model of Fuzhou University and Yuanxing Intelligent Drug Team was published in Nature sub-journal

Unexpected drug interactions (DDIs) are an important issue in drug research and clinical application because they are highly likely to cause serious adverse drug effects. reaction or drug withdrawal.
While many deep learning models have achieved good results in DDI prediction, model interpretability to reveal the underlying causes of DDI has not been widely explored.
Researchers from Fuzhou University, the First Affiliated Hospital of Fujian Medical University and Yuanxing Intelligent Medicine proposed MeTDDI - a deep learning framework with local-global self-attention and joint attention for learning based on DDI prediction plot of subject.
Regarding interpretability, researchers conducted an extensive evaluation on 73 drugs (13,786 DDIs), and MeTDDI can accurately explain the structural mechanisms of 5,602 DDIs involving 58 drugs. Furthermore, MeTDDI shows potential to explain complex DDI mechanisms and reduce DDI risk.
MeTDDI provides a new perspective for exploring DDI mechanisms, which will facilitate drug discovery and polypharmacy, thereby providing safer treatments for patients.
The study was titled "Learning motif-based graphs for drug–drug interaction prediction via local–global self-attention" and was published in "Nature Machine Intelligence" on August 27, 2024.

Due to aging and multimorbidity, drug combinations or polypharmacy are widely used and may have consequences for public health and the economy. Despite the therapeutic benefits of polypharmacy, there is a risk of unintended drug-drug interactions (DDIs), which may lead to serious adverse drug reactions (ADRs) or even discontinuation.
Thus, predicting DDIs in advance will bring huge benefits to drug research and clinical settings, thereby improving drug safety and protecting patient health. DDI assessment through in vitro and in vivo experiments is useful but is costly, time-consuming, and laborious, hindering the practicality of large-scale DDI screening.
Today, deep learning models have emerged as a promising alternative for high-throughput accurate DDI prediction as well as root cause explanation.
In the latest study, the research team of Fuzhou University, the First Affiliated Hospital of Fujian Medical University, and Yuanxing Intelligent Medicine focused on the prediction of metabolism-mediated drug interactions (MMDDI) and proposed a deep molecular structure-based Learning framework MeTDDI for predicting MMDDI.
This method is mainly used to solve three challenges in DDI prediction: (1) learning intra- and intermolecular substructural interactions, (2) predicting DDI-related drug metabolism, (3) widely providing and evaluating the model Interpretability.
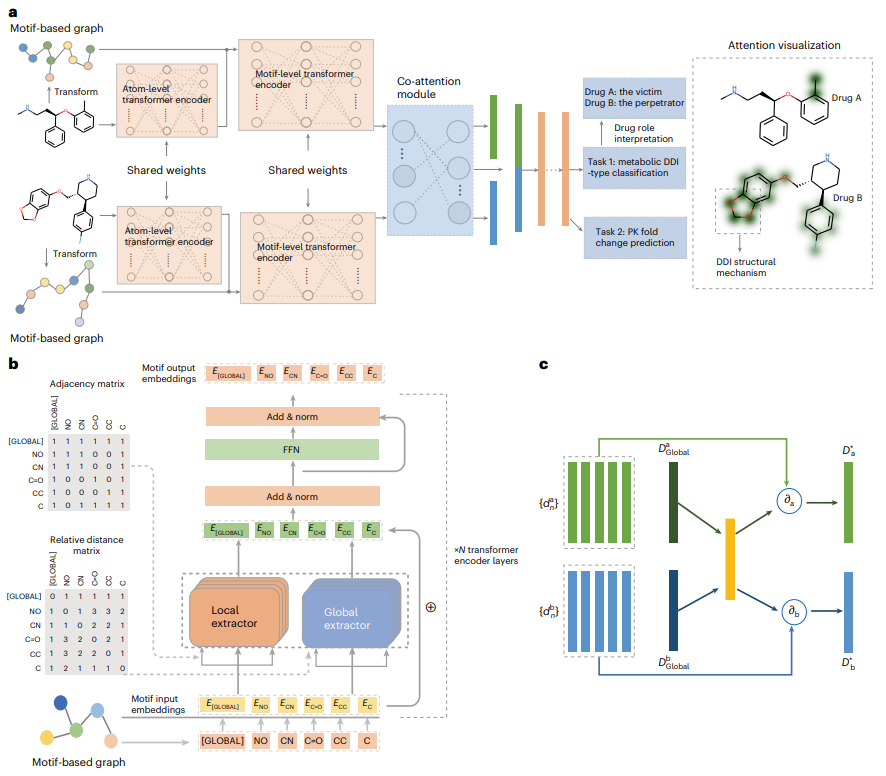
Benefiting from local-global self-attention and joint attention structures, MeTDDI can effectively learn intra- and intermolecular substructure interactions within/between graphs based on motifs , thereby performing DDI reasoning.
Evaluation results show that it achieves competitive performance compared to baselines in both classification and regression tasks. MeTDDI can also accurately identify the mechanistic role of a drug (perpetrator or victim) in DDI and quantify the impact of the perpetrator on the victim PK, which is very beneficial for both drug research and clinical applications.
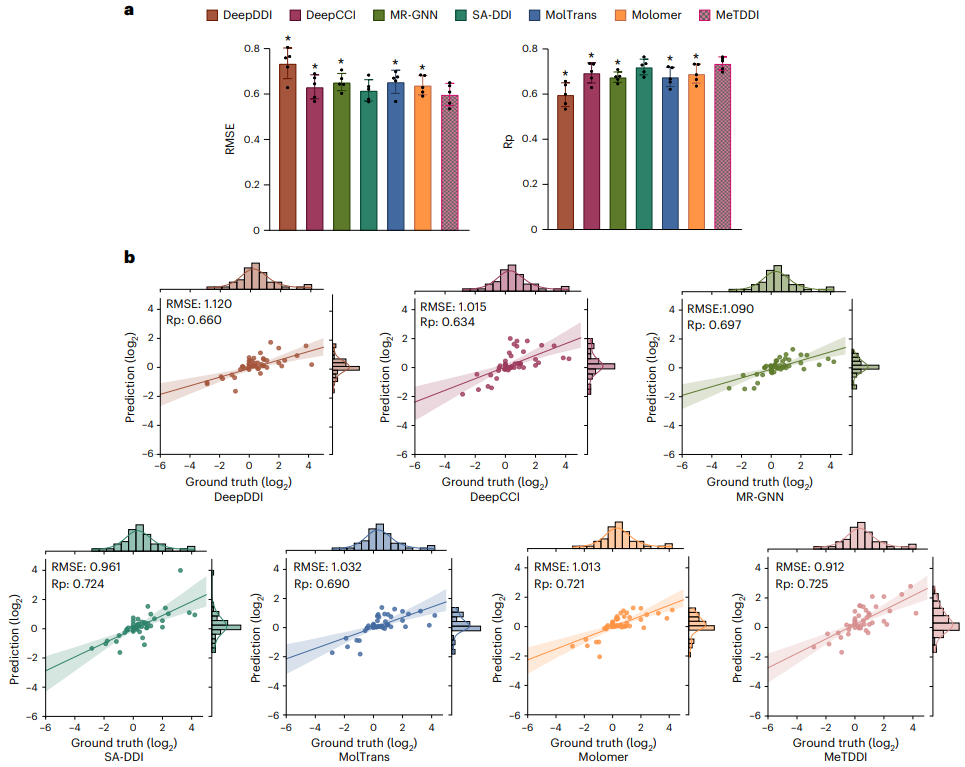
Regarding model interpretability, MeTDDI demonstrates the ability to identify key mechanistic substructures relevant to DDI.
First, the key substructures visualized by MeTDDI roughly match those reported in the literature from the analysis of 73 representative compounds (with 13,786 DDI pairs).
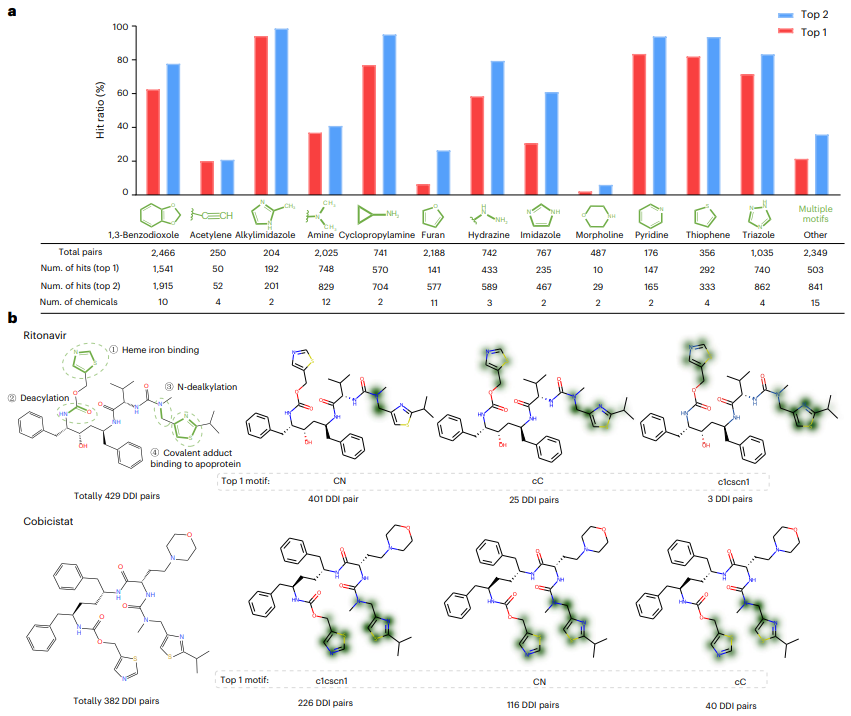
Second, the researchers evaluated the model interpretability of MeTDDI and two state-of-the-art models, namely CIGIN and CGIB. The results show that MeTDDI also exhibits excellent performance in terms of model interpretability.
Additionally, MeTDDI can highlight metabolic sites of chemicals associated with enzyme inhibition.
Advantages of MeTDDI
Traditional methods only explain the mechanism of DDI by testing the metabolic enzyme inhibition of the perpetrator in vitro, without fully considering the victim. This is problematic because the potency of enzyme inhibition by the perpetrator can vary depending on the chemical identity of the victim.
被害者は、代謝酵素 (特に CYP) と加害者の結合または相互作用パターンを変更し、その結果、さまざまな酵素阻害メカニズムが生じる可能性があります。これは、インビトロで単独で使用すると代謝酵素の強力な阻害剤であるエチニルエストラジオールやゲストデンなどの一部の化学物質が、それらの犠牲者と組み合わせると効果が低くなる理由を説明する可能性があります。これは、エチニルエストラジオールを用いた研究でなぜ 2 つの反応しか観察されなかったのかを説明する可能性があり、これが in vitro で CYP3A4 を不活化するメカニズムであると考えられています。
さらに、パロキセチンとイトラコナゾールのケーススタディでは、MeTDDI が化学物質のモチーフの変化を正確に予測し、生物学的実験の結果と一致していることが示されており、研究者が薬物の構造を変更して MMDDI のリスクを軽減するのに役立つ可能性が実証されています。
要約すると、MeTDDI は DDI 予測機能を強化し、DDI メカニズムを理解して探索するための新しい視点を提供します。これにより、医薬品開発とポリファーマシーが促進され、それによって患者により安全な治療が提供されます。
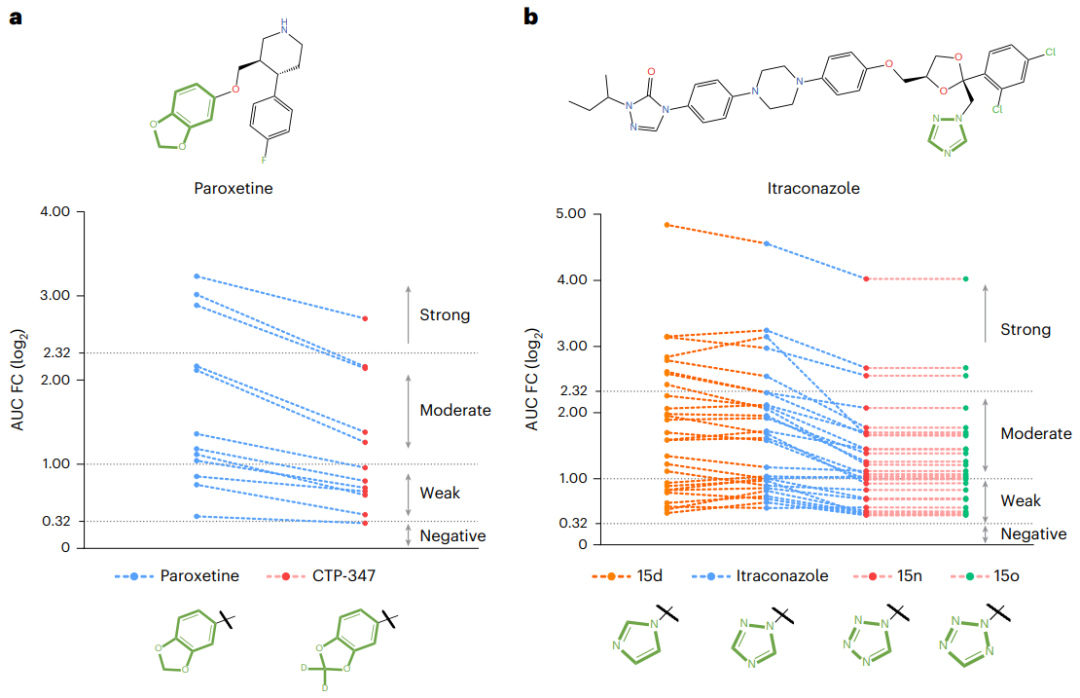
図: MeTDDI を使用した DDI 軽減の 2 つのケーススタディ。 (出典:論文) MeTDDI の改善方向
MeTDDI には多くの利点がありますが、同時にいくつかの制限もあります。
まず、困難なシナリオでは正確な予測が困難です。これは、DDI メカニズムの多様性と複雑さ、および薬物構造のみに依存することの限界に起因している可能性があります。
MMDDI では両方の薬物が同じ代謝酵素上で相互作用する必要があるため、酵素の特徴をモデルに組み込んで学習を改善できます。ただし、一部の代謝酵素 (CYP など) は薬物と酵素の相互作用部位に非常に高い柔軟性を示すため、酵素特性のモデル化は依然として課題です。
第二に、MeTDDI でトレーニングされたデータセットは FDA の医薬品ラベルに基づいています。これは集団の統計的観察であり、個々の患者の特徴を反映していない可能性があります。したがって、モデルを開発し、将来的により正確な予測を行うために、利用可能な場合は個々の患者データを考慮する必要があります。第三に、MeTDDI は 3 つ以上の薬物の相互作用を同時に予測することが難しい可能性があります。
ただし、ポリファーマシーを確保するための一般的な方法は、考えられるすべての薬物ペア間のペアごとの DDI を検索することです。MeTDDI を直接展開して、すべての薬物ペアを列挙することで複数の薬物間の DDI を予測できます。
最後に、DDI の基礎となる新たに発見された部分構造については、分子ドッキングなどの代替技術を補完的なアプローチとして採用して、MeTDDI 可視化機能の信頼性を高めることができます。そして研究者らは、分子ドッキングはMeTDDIを補完する貴重なツールであると述べている。
論文リンク: https://www.nature.com/articles/s42256-024-00888-6
The above is the detailed content of Efficiently and accurately predict DDI, the explanatory drug AI model of Fuzhou University and Yuanxing Intelligent Drug Team was published in Nature sub-journal. For more information, please follow other related articles on the PHP Chinese website!

Hot AI Tools
Undresser.AI Undress
AI-powered app for creating realistic nude photos
AI Clothes Remover
Online AI tool for removing clothes from photos.

Undress AI Tool
Undress images for free

Clothoff.io
AI clothes remover
Video Face Swap
Swap faces in any video effortlessly with our completely free AI face swap tool!

Hot Article

Hot Tools

Notepad++7.3.1
Easy-to-use and free code editor

SublimeText3 Chinese version
Chinese version, very easy to use

Zend Studio 13.0.1
Powerful PHP integrated development environment

Dreamweaver CS6
Visual web development tools

SublimeText3 Mac version
God-level code editing software (SublimeText3)

Hot Topics
 1664
1664
 14
14
 1423
1423
 52
52
 1317
1317
 25
25
 1268
1268
 29
29
 1246
1246
 24
24
 Breaking through the boundaries of traditional defect detection, 'Defect Spectrum' achieves ultra-high-precision and rich semantic industrial defect detection for the first time.
Jul 26, 2024 pm 05:38 PM
Breaking through the boundaries of traditional defect detection, 'Defect Spectrum' achieves ultra-high-precision and rich semantic industrial defect detection for the first time.
Jul 26, 2024 pm 05:38 PM
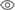
In modern manufacturing, accurate defect detection is not only the key to ensuring product quality, but also the core of improving production efficiency. However, existing defect detection datasets often lack the accuracy and semantic richness required for practical applications, resulting in models unable to identify specific defect categories or locations. In order to solve this problem, a top research team composed of Hong Kong University of Science and Technology Guangzhou and Simou Technology innovatively developed the "DefectSpectrum" data set, which provides detailed and semantically rich large-scale annotation of industrial defects. As shown in Table 1, compared with other industrial data sets, the "DefectSpectrum" data set provides the most defect annotations (5438 defect samples) and the most detailed defect classification (125 defect categories
 Training with millions of crystal data to solve the crystallographic phase problem, the deep learning method PhAI is published in Science
Aug 08, 2024 pm 09:22 PM
Training with millions of crystal data to solve the crystallographic phase problem, the deep learning method PhAI is published in Science
Aug 08, 2024 pm 09:22 PM
Editor |KX To this day, the structural detail and precision determined by crystallography, from simple metals to large membrane proteins, are unmatched by any other method. However, the biggest challenge, the so-called phase problem, remains retrieving phase information from experimentally determined amplitudes. Researchers at the University of Copenhagen in Denmark have developed a deep learning method called PhAI to solve crystal phase problems. A deep learning neural network trained using millions of artificial crystal structures and their corresponding synthetic diffraction data can generate accurate electron density maps. The study shows that this deep learning-based ab initio structural solution method can solve the phase problem at a resolution of only 2 Angstroms, which is equivalent to only 10% to 20% of the data available at atomic resolution, while traditional ab initio Calculation
 NVIDIA dialogue model ChatQA has evolved to version 2.0, with the context length mentioned at 128K
Jul 26, 2024 am 08:40 AM
NVIDIA dialogue model ChatQA has evolved to version 2.0, with the context length mentioned at 128K
Jul 26, 2024 am 08:40 AM
The open LLM community is an era when a hundred flowers bloom and compete. You can see Llama-3-70B-Instruct, QWen2-72B-Instruct, Nemotron-4-340B-Instruct, Mixtral-8x22BInstruct-v0.1 and many other excellent performers. Model. However, compared with proprietary large models represented by GPT-4-Turbo, open models still have significant gaps in many fields. In addition to general models, some open models that specialize in key areas have been developed, such as DeepSeek-Coder-V2 for programming and mathematics, and InternVL for visual-language tasks.
 Google AI won the IMO Mathematical Olympiad silver medal, the mathematical reasoning model AlphaProof was launched, and reinforcement learning is so back
Jul 26, 2024 pm 02:40 PM
Google AI won the IMO Mathematical Olympiad silver medal, the mathematical reasoning model AlphaProof was launched, and reinforcement learning is so back
Jul 26, 2024 pm 02:40 PM
For AI, Mathematical Olympiad is no longer a problem. On Thursday, Google DeepMind's artificial intelligence completed a feat: using AI to solve the real question of this year's International Mathematical Olympiad IMO, and it was just one step away from winning the gold medal. The IMO competition that just ended last week had six questions involving algebra, combinatorics, geometry and number theory. The hybrid AI system proposed by Google got four questions right and scored 28 points, reaching the silver medal level. Earlier this month, UCLA tenured professor Terence Tao had just promoted the AI Mathematical Olympiad (AIMO Progress Award) with a million-dollar prize. Unexpectedly, the level of AI problem solving had improved to this level before July. Do the questions simultaneously on IMO. The most difficult thing to do correctly is IMO, which has the longest history, the largest scale, and the most negative
 PRO | Why are large models based on MoE more worthy of attention?
Aug 07, 2024 pm 07:08 PM
PRO | Why are large models based on MoE more worthy of attention?
Aug 07, 2024 pm 07:08 PM
In 2023, almost every field of AI is evolving at an unprecedented speed. At the same time, AI is constantly pushing the technological boundaries of key tracks such as embodied intelligence and autonomous driving. Under the multi-modal trend, will the situation of Transformer as the mainstream architecture of AI large models be shaken? Why has exploring large models based on MoE (Mixed of Experts) architecture become a new trend in the industry? Can Large Vision Models (LVM) become a new breakthrough in general vision? ...From the 2023 PRO member newsletter of this site released in the past six months, we have selected 10 special interpretations that provide in-depth analysis of technological trends and industrial changes in the above fields to help you achieve your goals in the new year. be prepared. This interpretation comes from Week50 2023
 To provide a new scientific and complex question answering benchmark and evaluation system for large models, UNSW, Argonne, University of Chicago and other institutions jointly launched the SciQAG framework
Jul 25, 2024 am 06:42 AM
To provide a new scientific and complex question answering benchmark and evaluation system for large models, UNSW, Argonne, University of Chicago and other institutions jointly launched the SciQAG framework
Jul 25, 2024 am 06:42 AM
Editor |ScienceAI Question Answering (QA) data set plays a vital role in promoting natural language processing (NLP) research. High-quality QA data sets can not only be used to fine-tune models, but also effectively evaluate the capabilities of large language models (LLM), especially the ability to understand and reason about scientific knowledge. Although there are currently many scientific QA data sets covering medicine, chemistry, biology and other fields, these data sets still have some shortcomings. First, the data form is relatively simple, most of which are multiple-choice questions. They are easy to evaluate, but limit the model's answer selection range and cannot fully test the model's ability to answer scientific questions. In contrast, open-ended Q&A
 The accuracy rate reaches 60.8%. Zhejiang University's chemical retrosynthesis prediction model based on Transformer was published in the Nature sub-journal
Aug 06, 2024 pm 07:34 PM
The accuracy rate reaches 60.8%. Zhejiang University's chemical retrosynthesis prediction model based on Transformer was published in the Nature sub-journal
Aug 06, 2024 pm 07:34 PM
Editor | KX Retrosynthesis is a critical task in drug discovery and organic synthesis, and AI is increasingly used to speed up the process. Existing AI methods have unsatisfactory performance and limited diversity. In practice, chemical reactions often cause local molecular changes, with considerable overlap between reactants and products. Inspired by this, Hou Tingjun's team at Zhejiang University proposed to redefine single-step retrosynthetic prediction as a molecular string editing task, iteratively refining the target molecular string to generate precursor compounds. And an editing-based retrosynthetic model EditRetro is proposed, which can achieve high-quality and diverse predictions. Extensive experiments show that the model achieves excellent performance on the standard benchmark data set USPTO-50 K, with a top-1 accuracy of 60.8%.
 Nature's point of view: The testing of artificial intelligence in medicine is in chaos. What should be done?
Aug 22, 2024 pm 04:37 PM
Nature's point of view: The testing of artificial intelligence in medicine is in chaos. What should be done?
Aug 22, 2024 pm 04:37 PM
Editor | ScienceAI Based on limited clinical data, hundreds of medical algorithms have been approved. Scientists are debating who should test the tools and how best to do so. Devin Singh witnessed a pediatric patient in the emergency room suffer cardiac arrest while waiting for treatment for a long time, which prompted him to explore the application of AI to shorten wait times. Using triage data from SickKids emergency rooms, Singh and colleagues built a series of AI models that provide potential diagnoses and recommend tests. One study showed that these models can speed up doctor visits by 22.3%, speeding up the processing of results by nearly 3 hours per patient requiring a medical test. However, the success of artificial intelligence algorithms in research only verifies this